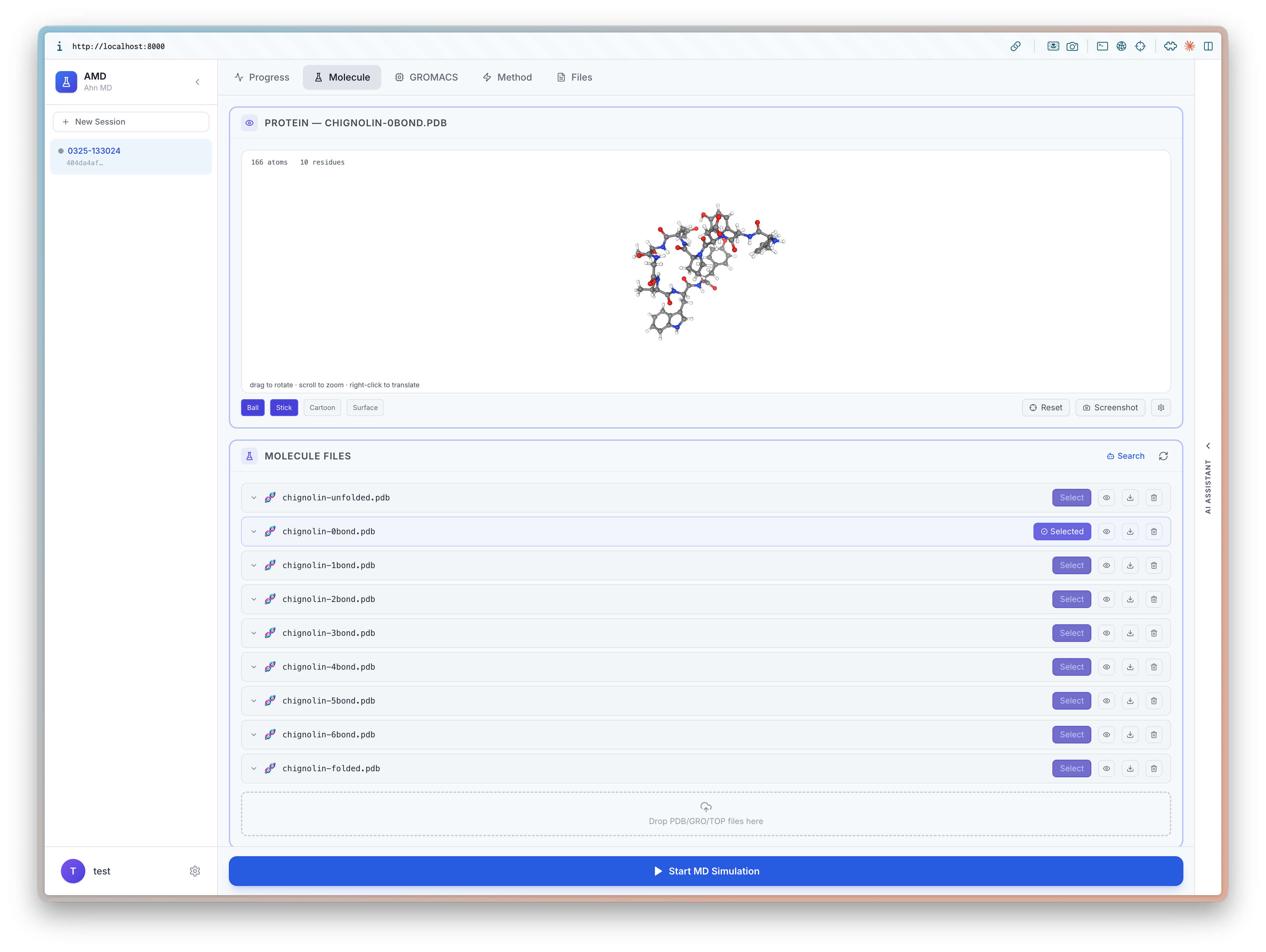
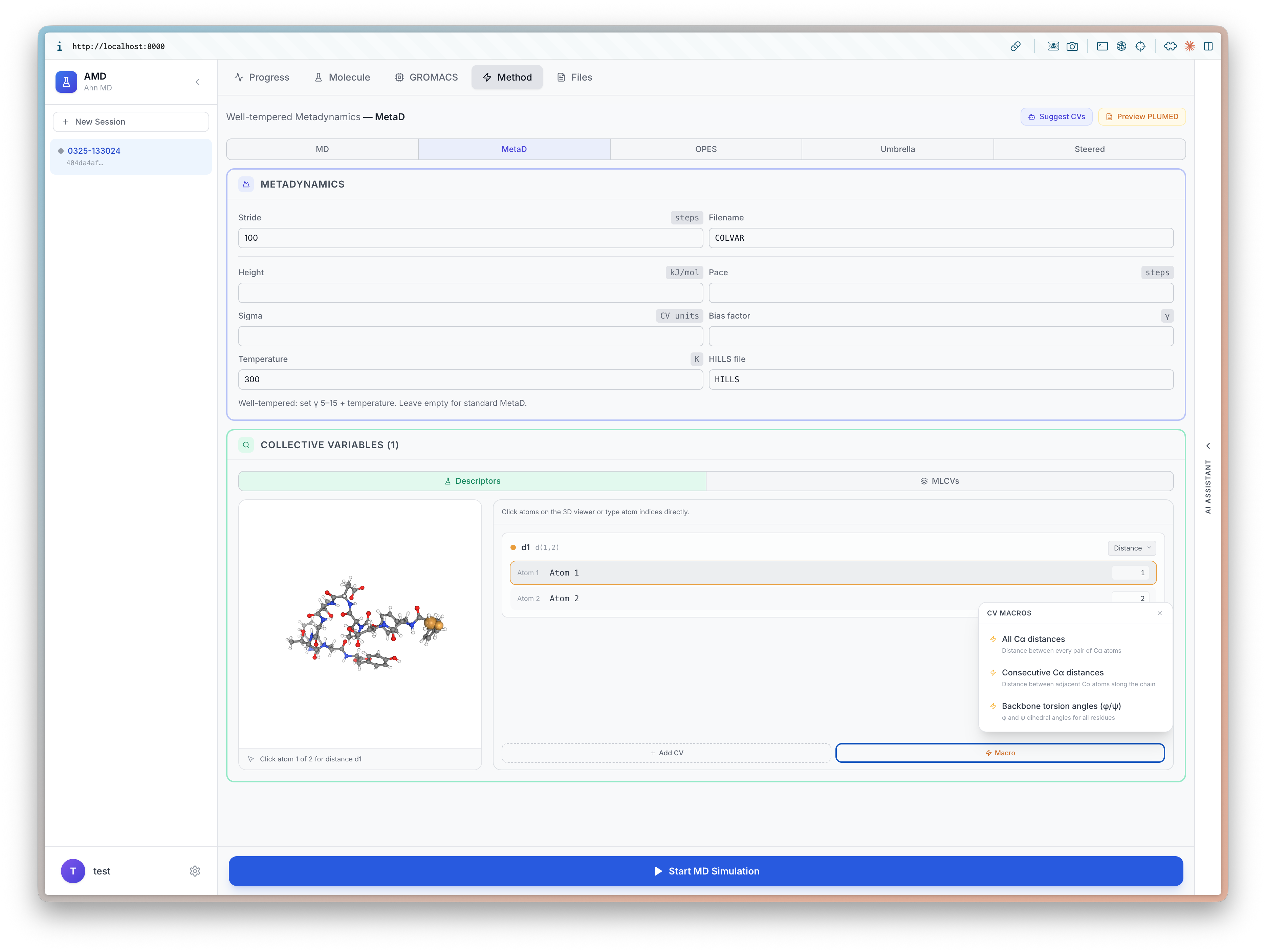
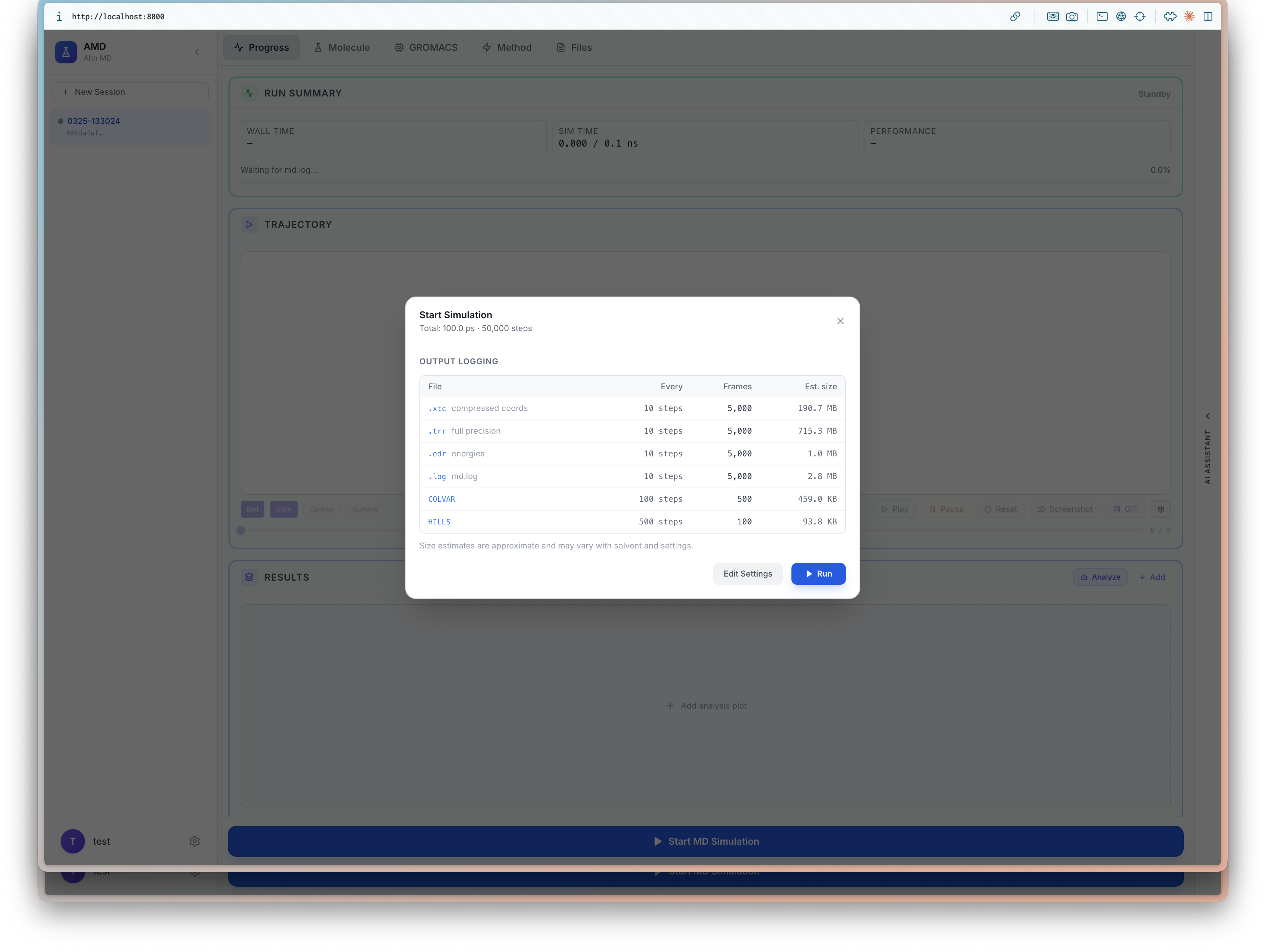
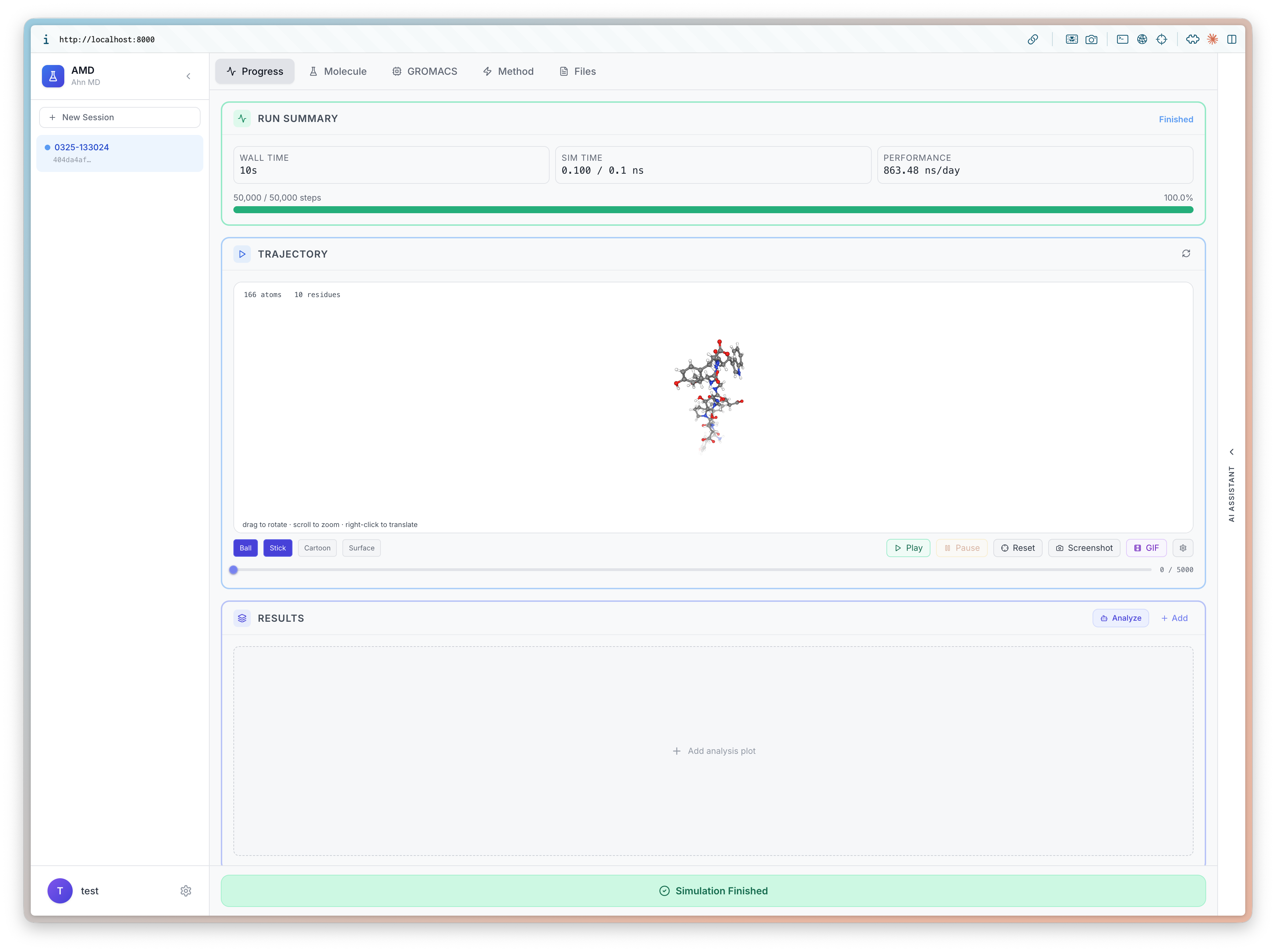
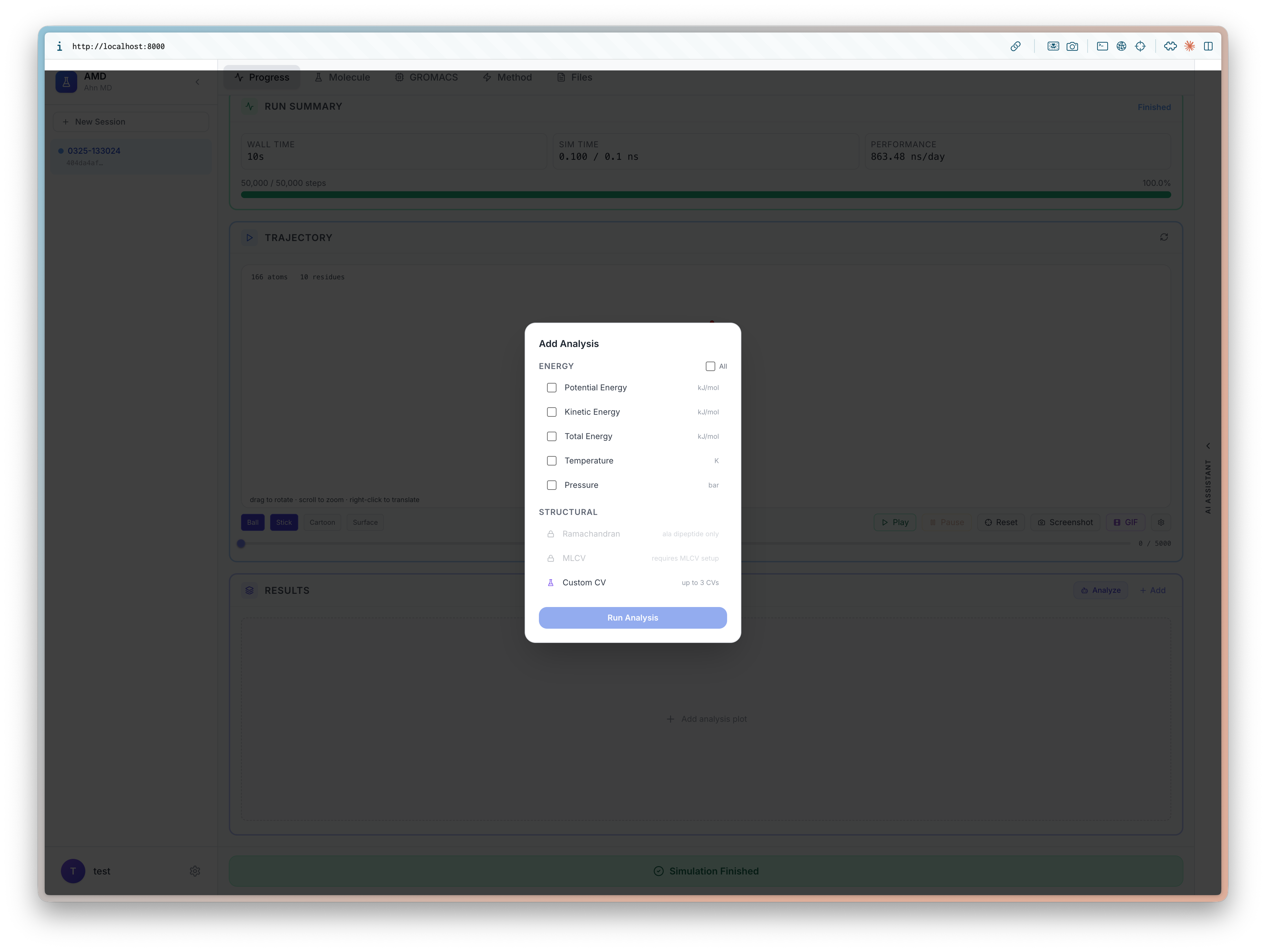
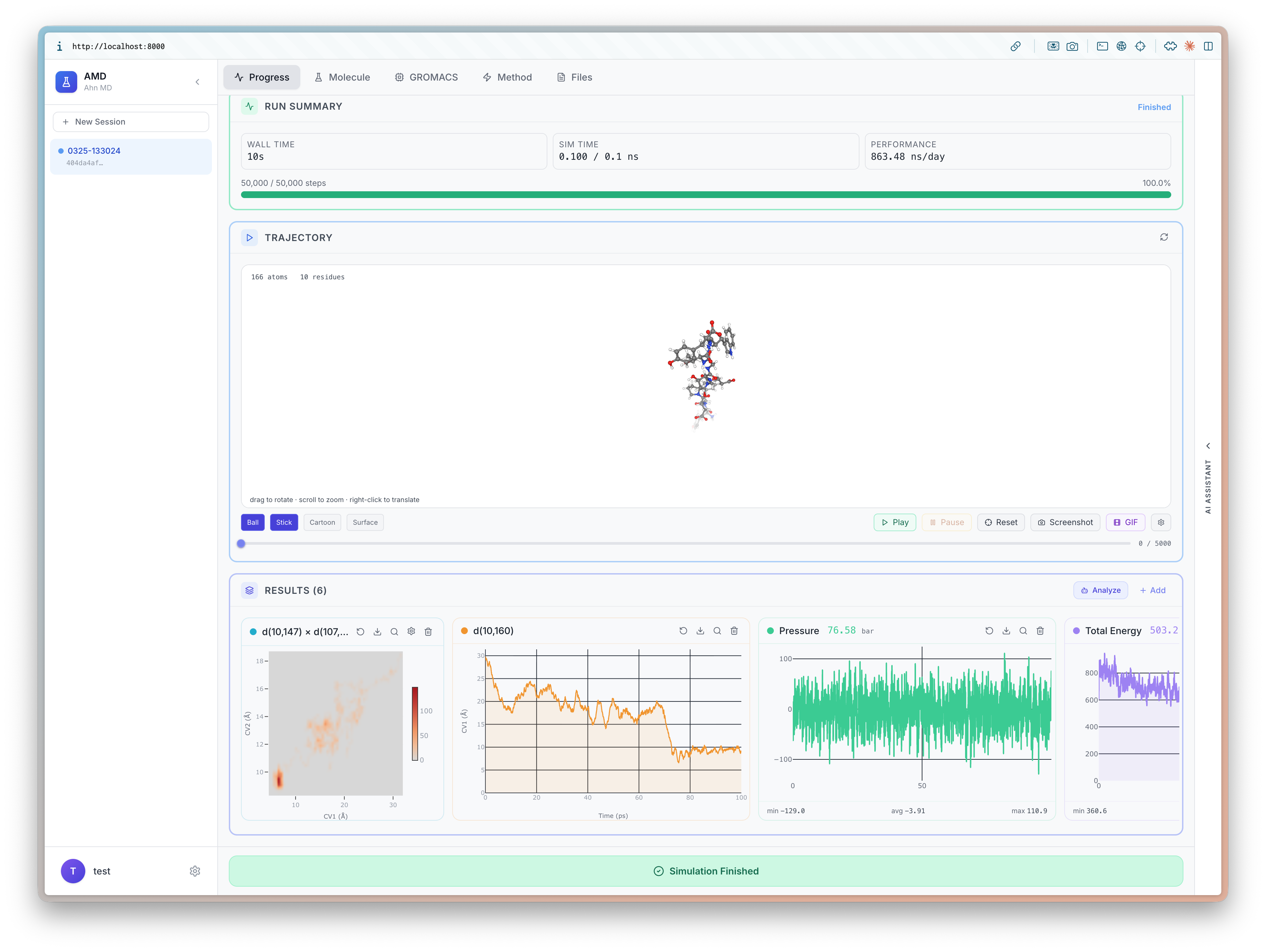
I will be at ICLR 2026 (4/23-27) to present BioEmu-CV See you in Rio 🇧🇷, I'd love to chat about small molecules and proteins!
Bio
Hello! I'm a 2nd year Ph.D. student at Kim Jaechul Graduate School of AI at KAIST, advised by Sungsoo Ahn. My research interests are mainly on AI4Science, integrating machine learning to advance scientific discovery on small molecules and proteins! I'm interested in the following topics:- Small molecule interactions (drug, ligand)
- Protein dynamics, Collective Variables (CVs)
- Graph Neural Networks (GNNs), Over-squashing
Publications
Most recent publications on Google Scholar.
† indicates equal contribution, ‡ indicates co corresponding author.
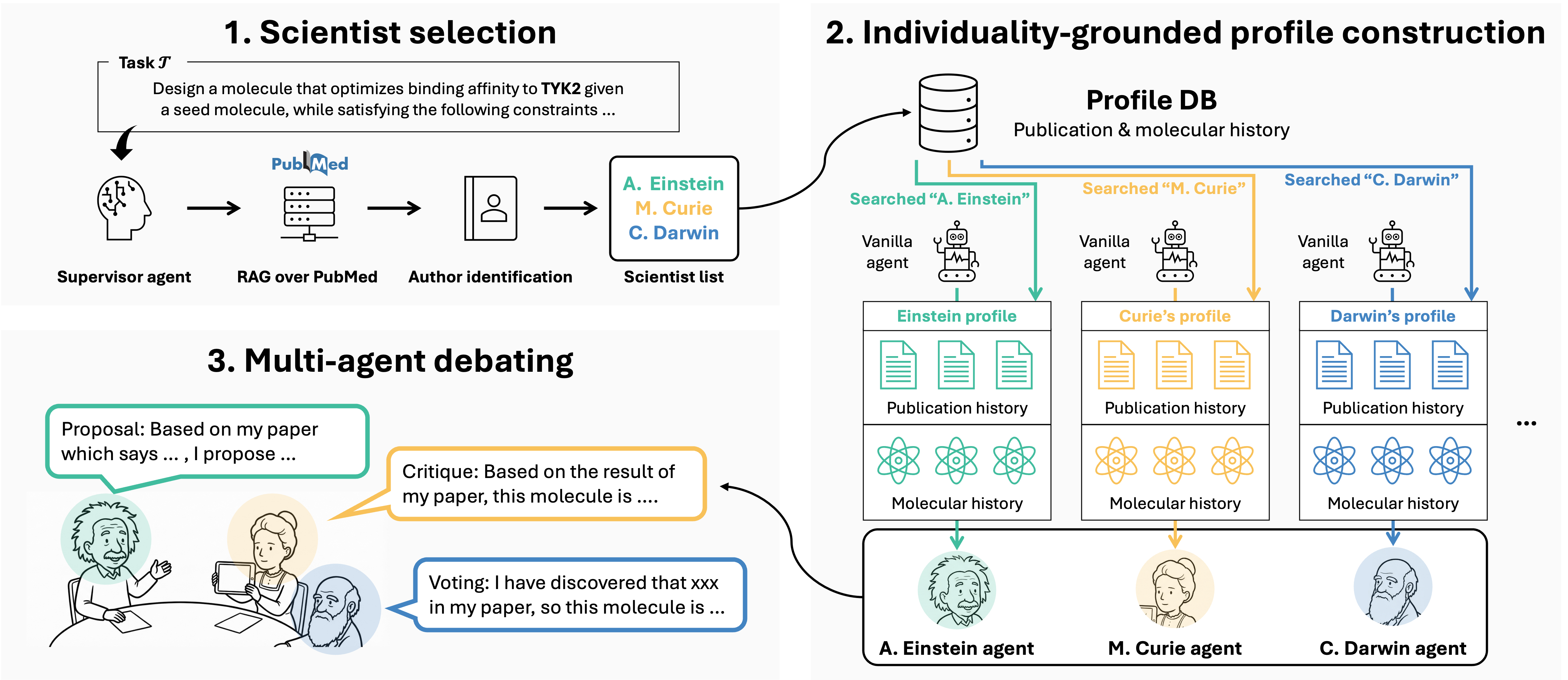
INDIBATOR: Diverse and Fact-Grounded Individuality for Multi-Agent Debate in Molecular Discovery
Yunhui Jang, Seonghyun Park, Jaehyung Kim, Sungsoo Ahn
Preprint, under review
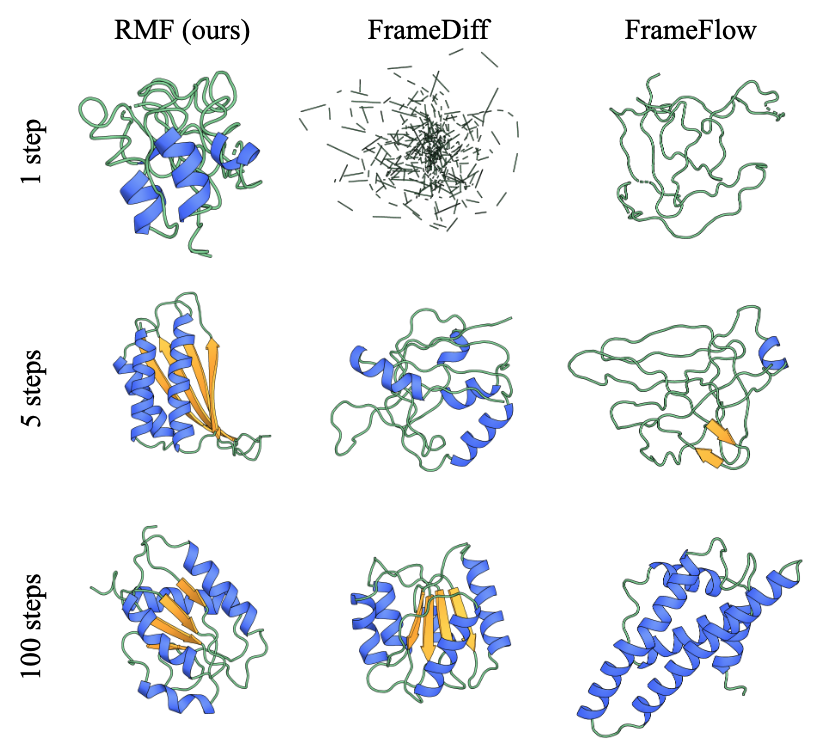
Dongyeop Woo, Marta Skreta, Seonghyun Park, Kirill Neklyudov, Sungsoo Ahn
Preprint, under review
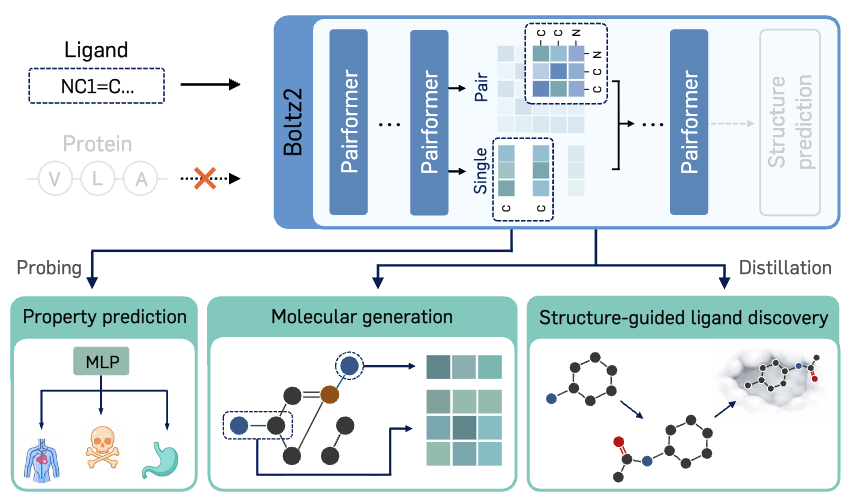
Boltz is a Strong Baseline for Atom-level Representation Learning
Hyosoon Jang, Hyunjin Seo, Yunhui Jang, Seonghyun Park, Sungsoo Ahn
Preprint, under review
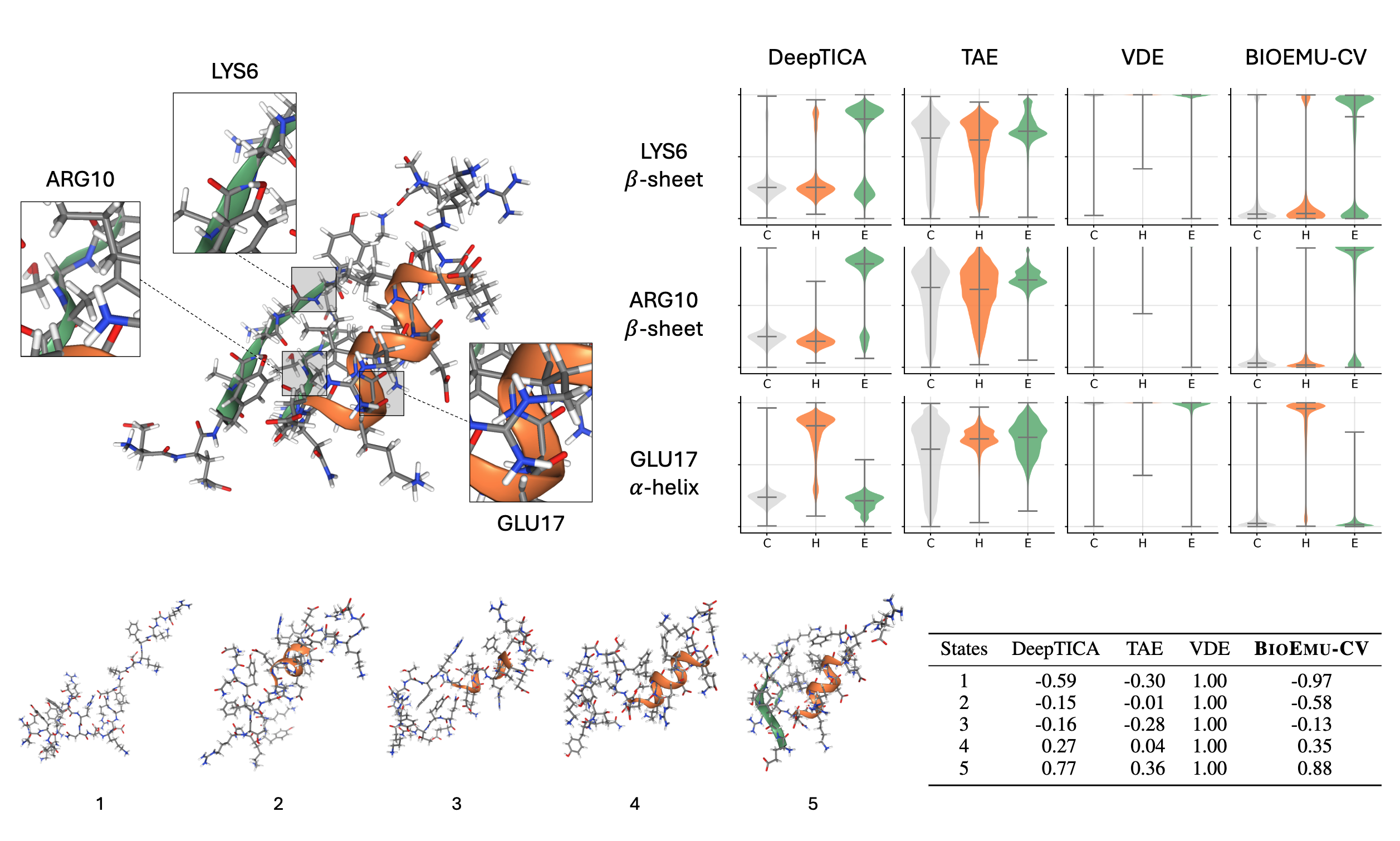
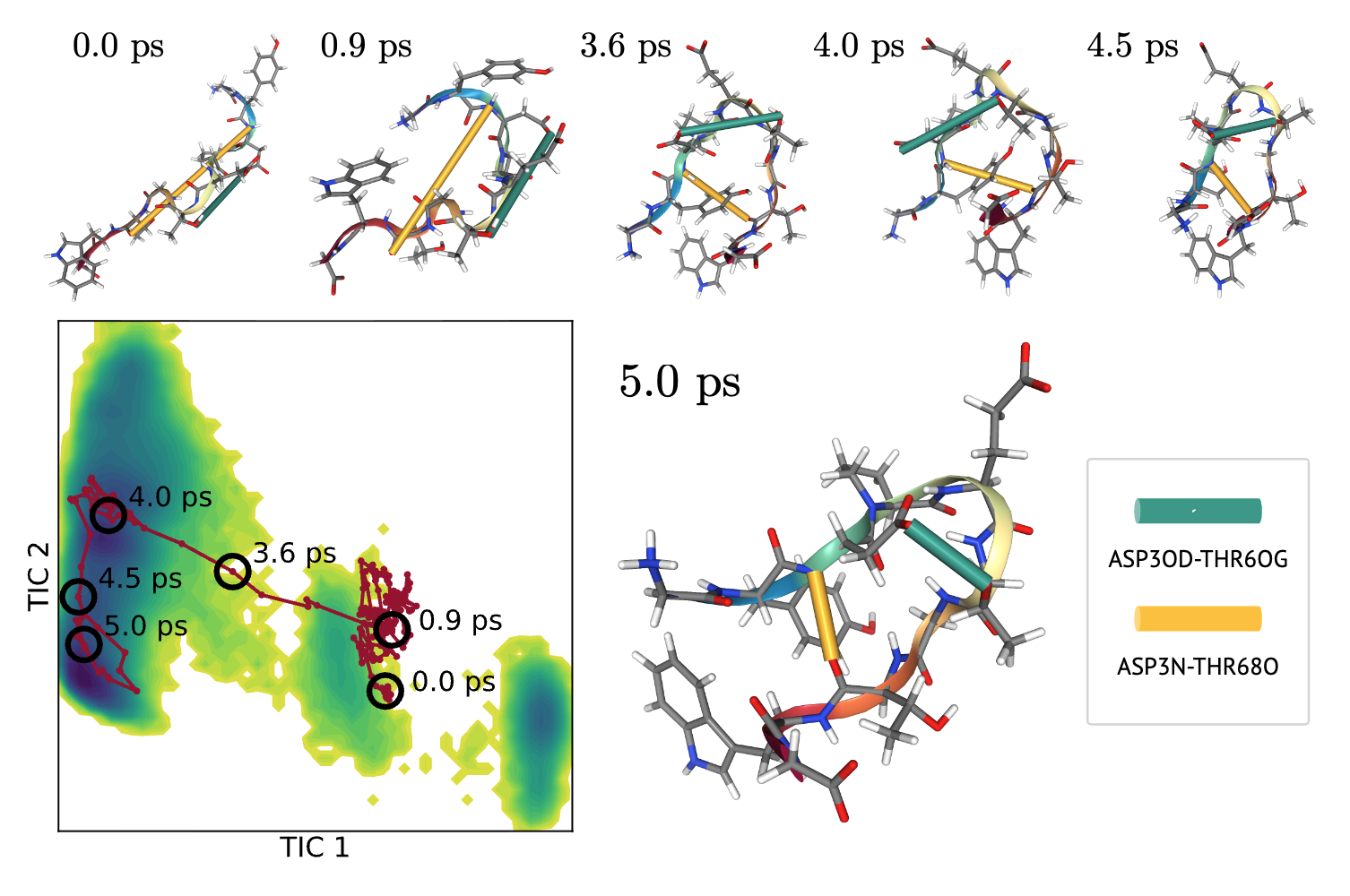
Learning Collective Variables from BioEmu with Time-lagged Generation
Seonghyun Park, Kiyoung Seong, Soojung Yang, Rafael Gómez-Bombarelli, Sungsoo Ahn
ICLR 2026
ICML 2025 Workshop, GenBio
Transition path sampling with improved off-policy training of diffusion path samplers
Kiyoung Seong, Seonghyun Park, Seonghwan Kim, Woo Youn Kim, Sungsoo Ahn
ICLR 2025
ICML 2024 Workshop, SPIGM
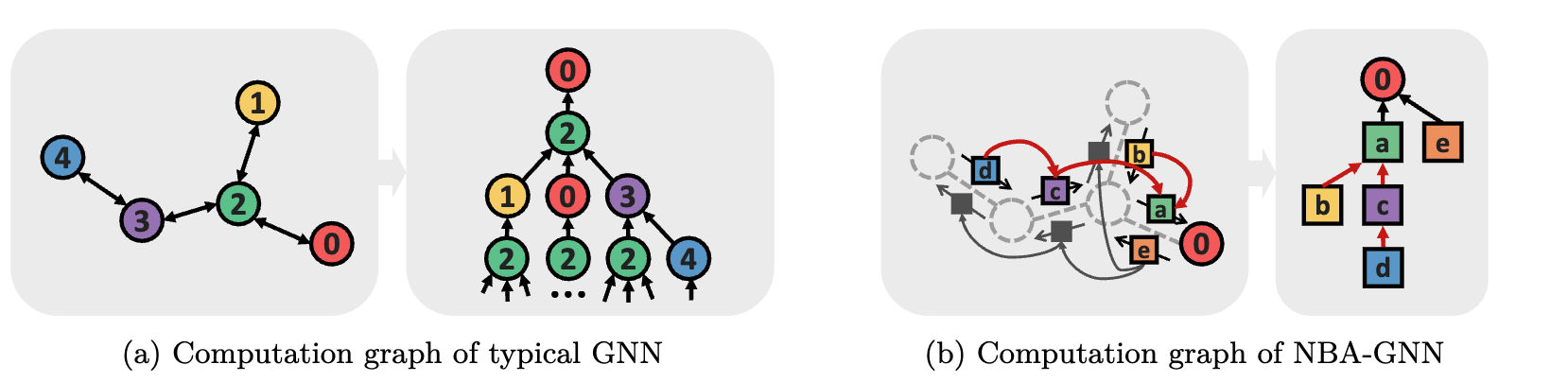
Non-backtracking Graph Neural Networks
Seonghyun Park†, Narae Ryu†, Gahee Kim, Dongyeop Woo, Se-Young Yun‡, Sungsoo Ahn‡
TMLR 2024
NeurIPS 2023 Workshop, New Frontiers in Graph Learning (GLFrontiers) (Oral)
INDIBATOR: Diverse and Fact-Grounded Individuality for Multi-Agent Debate in Molecular Discovery
Yunhui Jang, Seonghyun Park, Jaehyung Kim, Sungsoo Ahn
Preprint, under review
Dongyeop Woo, Marta Skreta, Seonghyun Park, Kirill Neklyudov, Sungsoo Ahn
Preprint, under review
Boltz is a Strong Baseline for Atom-level Representation Learning
Hyosoon Jang, Hyunjin Seo, Yunhui Jang, Seonghyun Park, Sungsoo Ahn
Preprint, under review
Learning Collective Variables from BioEmu with Time-lagged Generation
Seonghyun Park, Kiyoung Seong, Soojung Yang, Rafael Gómez-Bombarelli, Sungsoo Ahn
ICLR 2026
ICML 2025 Workshop, GenBio
Transition path sampling with improved off-policy training of diffusion path samplers
Kiyoung Seong, Seonghyun Park, Seonghwan Kim, Woo Youn Kim, Sungsoo Ahn
ICLR 2025
ICML 2024 Workshop, SPIGM
Non-backtracking Graph Neural Networks
Seonghyun Park†, Narae Ryu†, Gahee Kim, Dongyeop Woo, Se-Young Yun‡, Sungsoo Ahn‡
TMLR 2024
NeurIPS 2023 Workshop, New Frontiers in Graph Learning (GLFrontiers) (Oral)
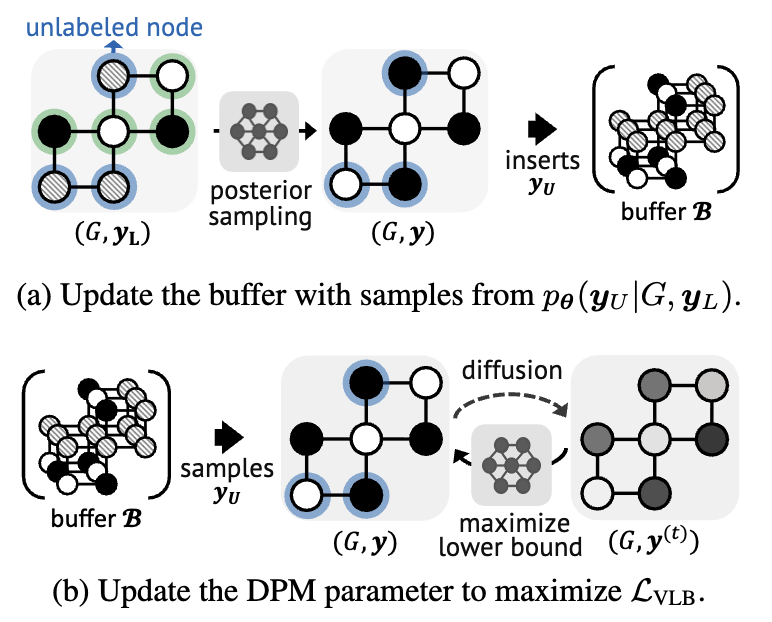
Diffusion Probabilistic Models for Structured Node Classification
Hyosoon Jang, Seonghyun Park, Sangwoo Mo, Sungsoo Ahn
NeurIPS 2023
ICML 2023 Workshop, Structured Probabilistic Inference & Generative Modeling
Side Projects
Vitæ
Full Resume in PDF.
-
KAIST 2026 -Technicial Research Personnel (Military Service)
Structured and Probabilistic Machine Learning (SPML) Lab -
KAIST AI 2025 -Ph.D. Student
Structured and Probabilistic Machine Learning (SPML) Lab -
POSTECH 2023 - 2025Master's Degree
Machine Learning (ML) Lab -
INSA Lyon Spring 2022Exchange student
Bioinformatique -
Bagelcode Summer 2021Buisness Analyst, Intern
BA Team -
Sellerhub Summer 2020Product Manager, Intern
PM Team -
POSTECH 2019 - 2022Bachelor's Degree
Computer Science and Engineering
Hobbies 🎶📷
↑ Click this!